from sklearn.linear_model import LinearRegression,SGDRegressor,Ridge,LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import mean_squared_error,classification_report
import numpy as np
import pandas as pd
def logistic():
column= ['Sample code number', 'Clump Thickness', 'Uniformity of Cell Size', 'Uniformity of Cell Shape',
'Marginal Adhesion','Single Epithelial Cell Size', 'Bare Nuclei', 'Bland Chromatin', 'Normal Nucleoli', 'Mitoses', 'Class']
data=pd.read_csv("https://archive.ics.uci.edu/ml/machine-learning-databases/breast-cancer-wisconsin/breast-cancer-wisconsin.data",names=column)
print(data)
data=data.replace(to_replace='?',value=np.nan)#缺失值进行处理
data=data.dropna()
x_train,x_test,y_train,y_test=train_test_split(data[column[1:10]],data[column[10]],test_size=0.25)
#进行标准化处理
std=StandardScaler()
x_train=std.fit_transform(x_train)
x_test=std.transform(x_test)
lg = LogisticRegression(C=1.0)
lg.fit(x_train, y_train)
y_predict=lg.predict(x_test)
print(lg.coef_)
print("准确率",lg.score(x_test,y_test))
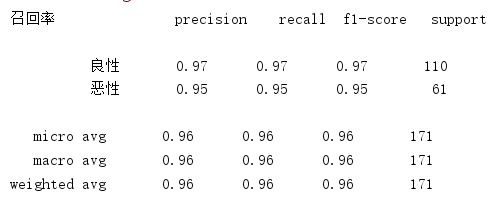
print("召回率",classification_report(y_test,y_predict,labels=[2,4],target_names=["良性","恶性"]))
return None
if __name__ =="__main__":
logistic()